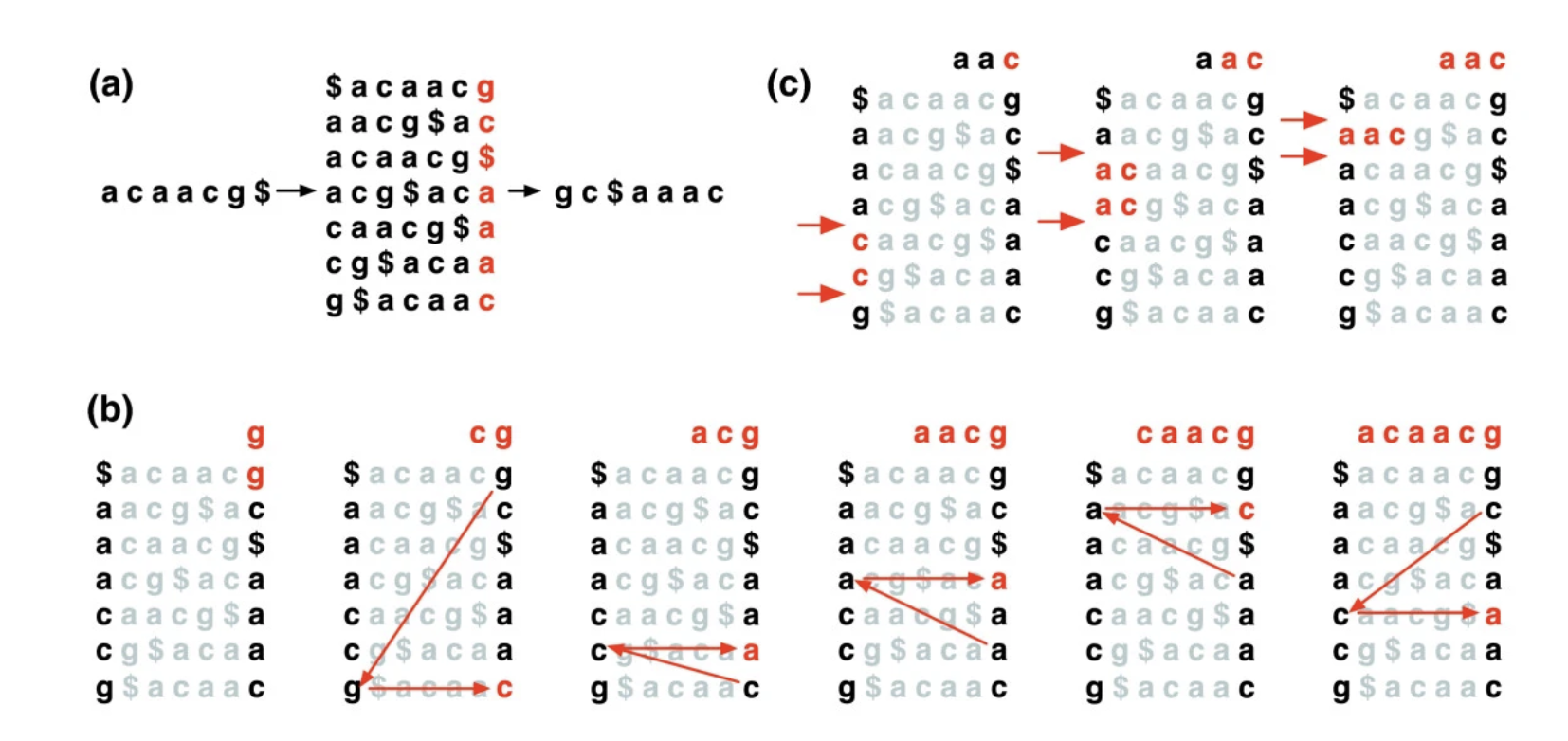
Dna Sequence Alignment With The Burrows Wheeler Transform
To name a few BurrowsWheeler transform is used in algorithms for sequence alignment image compression data compression etc. We implemented Burrows-Wheeler Alignment tool BWA a new read alignment package that is based on backward search with. We designed and implemented a new algorithm Burrows-Wheeler Aligners Smith-Waterman Alignment BWA-SW to align long. The Burrows-Wheeler Transform is a string compression technique that is used in compression tools such as bzip2 Using the FM-index 4 a data. We implemented Burrows-Wheeler Alignment tool BWA a new read alignment package that is based on backward search with Burrows..
Im writing Burrows-Wheeler Transform and its reverse in Python It works fine for small strings. The Burrows Wheeler Transform BWT was developed in 1994 by Michael Burrows and David..
To name a few BurrowsWheeler transform is used in algorithms for sequence alignment image compression data compression etc. We implemented Burrows-Wheeler Alignment tool BWA a new read alignment package that is based on backward search with. We designed and implemented a new algorithm Burrows-Wheeler Aligners Smith-Waterman Alignment BWA-SW to align long. The Burrows-Wheeler Transform is a string compression technique that is used in compression tools such as bzip2 Using the FM-index 4 a data. We implemented Burrows-Wheeler Alignment tool BWA a new read alignment package that is based on backward search with Burrows..
Introduction BWA is a software package for mapping low-divergent sequences against a large. Introduction to Burrows-Wheeler Alignment bwa and Samtools for..

Comments